

So, why do we need to use GNN?
The main reason is that we can't convert a graph to an N-dimensional vector or a sequence of them – that's why we can't use more straightforward approaches and neural network architectures to deal with such type of data.
What types of task can GNN perform?
Let's give an example of some problems that GNN can solve:
Node classification. In node classification, the task is to predict the node type for all the nodes in a graph. This type of problem is usually trained in a semi-supervised way, where only part of the graph is labeled. Typical applications for node classification include citation networks, social network posts and users classification. Link prediction. In link prediction, the task is to understand the relationship between entities in graphs and predict if two entities have a connection in between. For example, a recommender system can be treated as a link prediction problem where the model is given a set of users' reviews of different products, the task is to predict the users' preferences and tune the recommender system to push more relevant products according to users' interest. Representation learning. During the GNN training, most of the architectures involve the outputs that produce node embeddings that contain both structural information from the graph regarding the specific nodes and node feature descriptions. These outputs can be further used as the input for other models or components of the initial model (e.g. be sent to the multilayer perceptron for classification). Graph classification. The task is to determine the type or class of the whole graph into different categories. For example, we can try to classify whether a specific molecule (which structure is represented by the graph) has a useful property in biomedical or chemistry spheres.
What types of GNN exist?
Recurrent Graph Neural Network
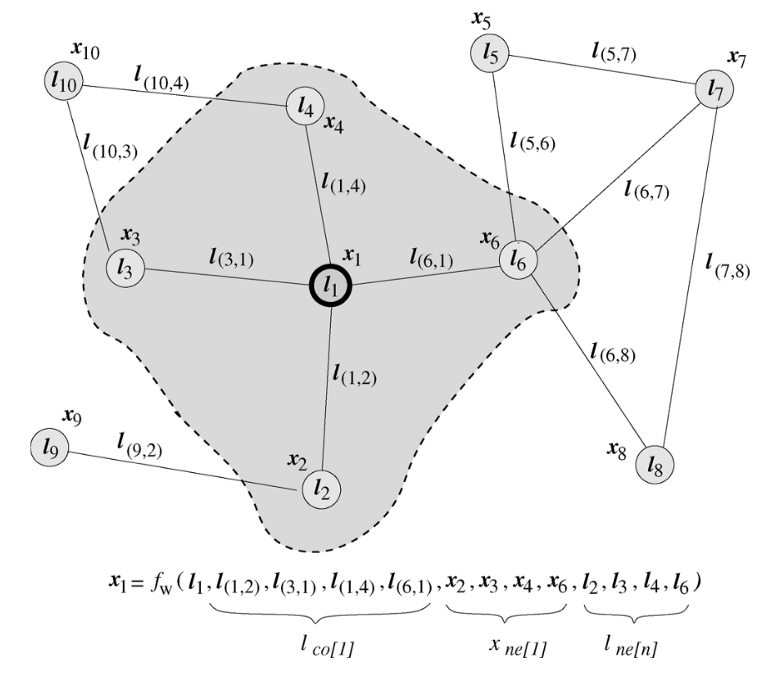
Recurrent Graph Neural Network – it's the first designed GNN architecture introduced in the original GNN paper. Its main idea is connected with iterative update of the node "state" – the computed function value utilizing the information about node neighborhood states.
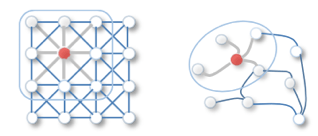
The idea of convolution on a graph is almost similar as in image convolution. In case of working with image, we sum the neighboring pixels around a center pixel, specified by a filter with parameterized size and learnable weight. Spatial Convolutional Network adopts the same idea by aggregating the features of neighboring nodes into the center node.
Recommender systems
Many companies use graph neural networks to build recommender systems. Typically, graphs are used to model user interaction with products and learn embeddings based on a set of properly selected negative samples. By ranking the results, personalized product offers are selected and shown to specific users in real time. One of the first services with such mechanism was Uber Eats – the GraphSage neural network selects food and restaurant recommendations.
Although the graphs are relatively small in the case of food recommendations, some companies use neural networks with billions of connections. For example, Alibaba has launched graph embeddings and graph neural networks for billions of users and products. The mere creation of such graphs is a nightmare for developers. Thanks to the Aligraph pipeline, you can build a graph with 400 million nodes in just five minutes. Aligraph supports efficient, distributed graph storage, optimized fetch operators, and a bunch of native graph neural networks. This pipeline is now used for recommendations and personalized searches across the company's many products.
Computer vision
Objects in the real world are deeply interconnected, so images of these objects can be successfully processed using graph neural networks. For example, you can perceive the content of an image through scene graphs – a set of objects in a picture with their relationships. Scene graphs are used to find images, understand and comprehend their content, add subtitles, answer visual questions, and generate images. These graphs can greatly improve the performance of models.
In one of the works of Facebook it is described that you can put objects from the popular COCO dataset into the frame, set their positions and sizes, and based on this information a scene graph will be created. With its help, the graph neural network determines the embeddings of objects, from which, in turn, the convolutional neural network creates object masks, frames and contours. End users can simply add new nodes to the graph (determining the relative position and size of nodes) so that neural networks can generate images with these objects.
Physics and chemistry
Representing the interactions between particles or molecules in the form of graphs and predicting the properties of new materials and substances using graph neural networks allows solving various natural science problems. For example, as part of the Open Catalyst project, Facebook and CMU are looking for new ways to store renewable energy from the sun and wind. One possible solution is to convert this energy through chemical reactions into other fuels, such as hydrogen. But for this, it is necessary to create new catalysts for high-intensity chemical reactions, and the methods known today like DFT are very expensive. The authors of the project posted the largest selection of catalysts and base layers for graph neural networks. The developers hope to find new low-cost molecular simulations that will complement the current expensive simulations that run in days with efficient energy and intermolecular force estimates that are computed in milliseconds.
Researchers at DeepMind have also used graph neural networks to emulate the dynamics of complex particle systems such as water and sand. By predicting at each step the relative motion of each particle, one can plausibly recreate the dynamics of the entire system and learn more about the laws that govern this motion. For example, this is how they try to solve the most interesting of the unsolved problems in the theory of solids – the transition to a glassy state. Graph neural networks not only allow you to emulate the dynamics during the transition, but also help you better understand how particles affect each other depending on time and distance.
Drug development
Pharmaceutical companies are actively looking for new ways to develop drugs, competing fiercely with each other and spending billions of dollars on research. In biology, you can use graphs to represent interactions at different levels. For example, at the molecular level, bonds between nodes would represent interatomic forces in a molecule, or interactions between amino acid bases in a protein. On a larger scale, graphs can represent interactions between proteins and RNA or metabolic products. Depending on the level of abstraction, graphs can be used for target identification, molecular property prediction, high-throughput screening, drug design, protein engineering, and drug repurposing.
Perhaps the most promising result of the use of graph neural networks in this area was the work of researchers from MIT, published in Cell in 2020. They applied a deep learning model called Chemprop , which predicted the antibiotic properties of the molecules: inhibition of E. coli reproduction. After training on just 2,500 molecules from a FDA-approved library, Chemprop was applied to a larger dataset, including a Drug Repurposing Hub containing the Halicin molecule. It is noteworthy that until now, Halicin has only been studied in relation to the treatment of diabetes, because its structure is very different from known antibiotics. But clinical experiments in vitro and in vivo have shown that Halicin is a broad spectrum antibiotic. Extensive comparison with strong neural network models highlighted the importance of Halicin's properties discovered using graph neural networks. In addition to the practical role of this work, the Chemprop architecture is also interesting for others: unlike many graph neural networks, it contains 5 layers and 1600 hidden dimensions, which is much more than the typical parameters of graph neural networks for such tasks. It can be just one of the few AI discoveries in the future new medicine.
